OncoLUNG Dx – Lung Cancer Diagnosis Test
Overview
OncoLUNG Dx is an innovative, non-invasive, accurate, and cost-effective already validated solution to help in Lung Cancer diagnosis as well as confirmatory diagnostic ―as an adjunct to suspicious image procedures findings, in order to reduce the number of unnecessary tissue biopsies that patients have to undergo―. It also has potentially uses for screening, prognosis and recurrence monitoring.
OncoLUNG Dx is based on a score calculation that it is obtained from several Biomarkers of the patient (mainly Tumor Markers but also patient’s clinical information).

Click here to download the brochure in PDF format.
Tumor Markers
Tumor Markers are parameters released by tumor cells, which enter the bloodstream or other biological fluids and are useful for the diagnosis, prognosis and treatment monitoring.
Most Tumor Markers are not specific to any type of cancer and the differences between benign and malignant diseases are quantitative (for example, patients with epithelial tumors tend to have significantly higher levels of these Tumor Markers than patients without malignancy).
There are now more than 20 well known parameters that are widely regarded as Tumor Markers such as PSA ―related to Prostate Cancer―, CA 15.3 ―related to Breast Cancer―, CA 125 and HE4 ―both related to Ovarian Cancer―, CEA and CA 19.9 ―both related to different gastrointestinal cancers (Colorectal, Gastric and Pancreatic Cancer)―, or NSE and ProGRP ―both related to in Lung Cancer―.
However, there are a variety of factors that can affect the accuracy of Tumor Markers by increasing its levels without malignancy presence. The main reason are benign diseases, among others, such as technical interferences.
In this sense, the Spanish Society of Clinical Biochemistry and Molecular Pathology, Cancer Biomarker Commission established the Barcelona Criteria, 4 criteria that help to correctly distinguish and value Tumor Markers results and reduce False Positives (FP):
- Tumor Markers Serum concentrations.
- Discard benign pathology by the exclusion of main source False Positive results.
- Follow-up if Tumor Markers moderate results (Grey Zone/Undetermined).
- Technical interference.
Statistical measurements in diagnostic tests
Unfortunately, the use of Tumor Markers in routine presents also other problems such as low Sensitivity in early stages, or nonexistence of any specific Tumor Marker for each malignant tumor. However, the combination of 2 or more Tumor Markers has a better outcome, especially in advanced stages.
In this regard, the combination of several Tumor Markers ―as well as the inclusion of patient history information in the equations―, using complex algorithms with multiple variables, results in higher Sensitivity and Specificity: that is what we have christened Multi-Biomarker Disease Activity Algorithm (MBDAA).
The Sensitivity of a diagnostic test is the percentage of actual positives that are correctly identified, and Specificity is the proportion of true negatives that are correctly classified. Both variables are closely linked together and give an idea of the accuracy of a test.
A test that correctly identifies all true positive as positive, but has many false negatives would have a Sensitivity of 100%, but low Specificity. For example, Sensitivity measures the number of cancerous tumors that are correctly identified as cancerous, whereas Specificity measures how many benign tumors are correctly identified as benign. A high Sensitivity means fewer cancers diagnosed as benign and high Specificity means fewer benign tumors diagnosed as cancerous.
Besides, the positive predictive value (PPV) is the number of true positives correctly identified on total real positive. A test with many false positives will have a low VPP. Moreover, the negative predictive value (NPV) is the number of true negatives correctly identified on the total actual negative. A high NPV value means that very few true positives were incorrectly identified as negative.
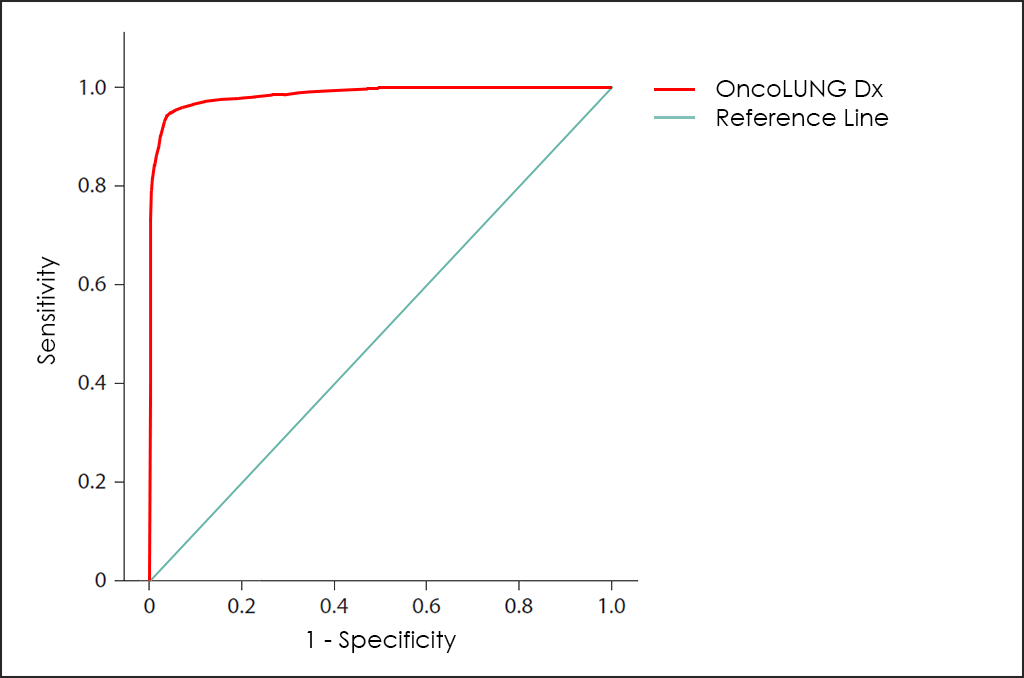
All these different values can be plotted together in a graphic that it is known as Receiving Operator Curve (ROC), where better results are displayed with curves that tends to come near to the upper left corner of the image (where 100% Sensitivity and 100% Specificity are reached).
Receiving Operator Curves (ROC)
The ROC curve of OncoLUNG Dx test ―based on the combined count of CA 15.3, CEA, CYFRA 21-1, NSE, SCC and ProGRP Tumor Markers; comorbidities; and other data from 4.296 consecutive patients, then fine-tuned by other research―, throws really interesting diagnostic capabilities: 93.5% Sensitivity and 96.2% Specificity.
Besides Risk: Histologycal Type and Subtype
OncoLUNG Dx test also allow us to determine with that so high accuracy the Histologycal Type, distinguishing between Non-Small Cell Lung Cancer (NSCLC) or Small Cell Lung Cancer (SCLC) as well as the Subtype, discriminating between Adenocarcinoma or Squamous Cell Carcinoma (if NSCLC Type).
How does it work
As all BIOPROGNOS’ MBDAA tests, OncoLUNG Dx test is available online once access is granted through our secure Cloud Platform. As a Cloud solution, it is designed to be used in a Software as a Service (SaaS) basis, that means, no installation, no periodically patch upgrades, low TCO (Total Cost of Ownership) and no maintenance.
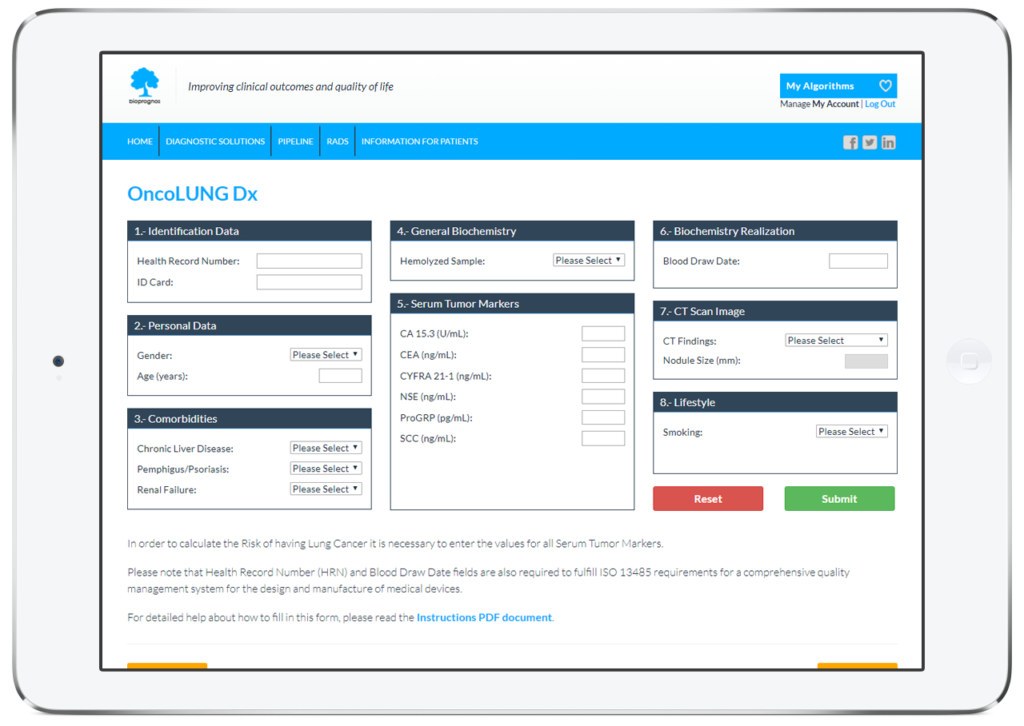
In this way, doctors or lab technicians only should fill the form with values obtained previously from patients (personal data, comorbidities, biochemistry values, CT Scan finding and lifestyle information), and click on Submit button in order to obtain the risk score of having Lung Cancer.
Order Form
To facilitate work, doctors can download and fill the Order Form for OncoLUNG Dx in a quick and an easy way ―with all required data for a best risk calculation already detailed―.
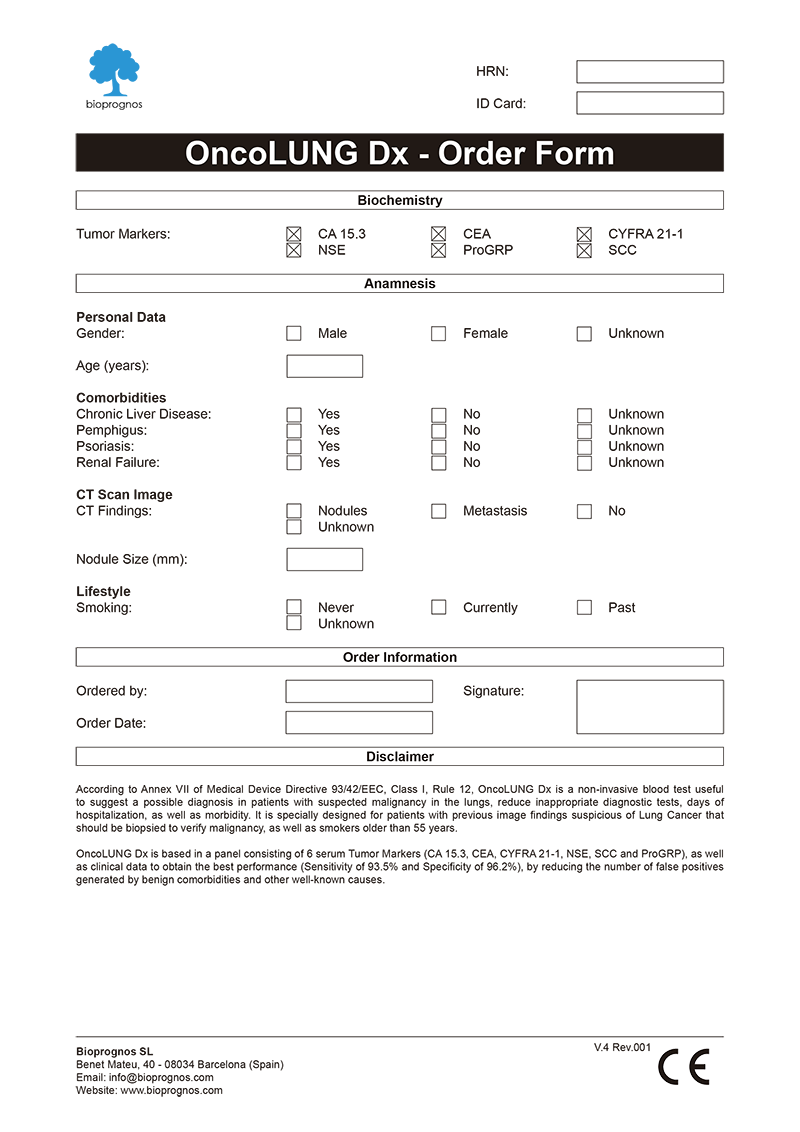

Click here to download the OncoLUNG Dx Order Form in PDF format.
Final Report
After doctors entered the patient’s data, OncoLUNG Dx test presents the results in a separate screen that can be converted to a PDF document in order to be downloaded or sent by email.
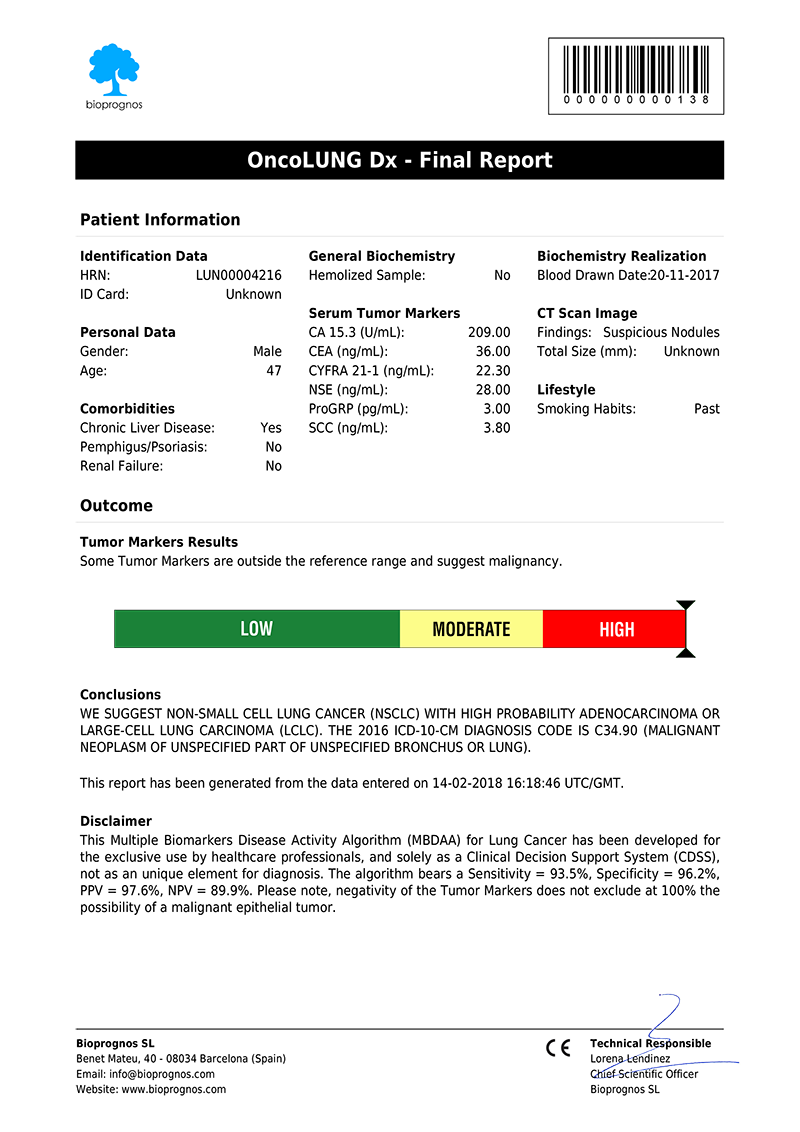

Click here to download the report in PDF format.
The report includes two main sections: Patient Information and Outcome. In the first one, all patient data entered previously is showed as record. The second one includes: Results, indicating whether Tumor Marker levels are within normal range or not; Risk, with a score bar showing the probability of having Lung Cancer; Comments, that are created dynamically to help doctors and healthcare professionals to understand ―in an easy way―, how to detect False Positives (FP), such as levels of Tumor Markers that would suspect the presence of Cancer, but when considering other variables together ―Comorbidities or Smoking Habits―, do not correspond with malignant diseases; and finally, Conclusions, with recommendations suggesting to retest patient in 1 year (for Low Risk), or in 4 weeks (for Moderate Risk, that is, these cases in which Tumor Marker levels are higher than normality but there is not quite clear to be High Risk.
Please note that final report is oriented to healthcare professionals only ―not to patients―, because it was designed as “a tool to help healthcare professionals in Lung Cancer diagnostic”, and also certified by obtaining the CE DECLARATION OF CONFORMITY (Medical Device Directive 93/42/EEC, Class I, rule 12).
CE Declaration of Conformity
Since November 22th, 2016 (when version 1.0 was released), OncoLUNG Dx test has the CE Declaration of Conformity that certifies it has been assessed to meet high safety, health, and environmental protection requirements.

Click here to download the OncoLUNG Dx CE DECLARATION OF CONFORMITY in PDF format.
This declaration also certifies that OncoLUNG Dx test can be sold throughout the European Economic Area (EEA) without restrictions.
Besides, there are two main benefits CE marking brings to businesses and consumers within the EEA:
- Businesses know that products bearing the CE marking can be traded in the EEA.
- Consumers enjoy the same level of health, safety, and environmental protection throughout the entire EEA.
Other tests in the market
20/20 GeneSystems, an American biotech company, have developed the PAULA’s (Protein Assays Utilizing Lung Cancer Analytes) Test, a multiplex immunoassay blood test that incorporates both tumor antigens and autoantibodies to determine the risk that Non-Small Cell Lung Cancer (NSCLC) is present in individuals from a high-risk population.
PAULA’s Test is based in 4 biomarkers found in serum, such as CA 125, CEA, CYFRA and NY-ESO-1, and the global performance is quite poor, as you can see in superposed ROC curves for PAULA Test and our OncoLUNG Dx Test:
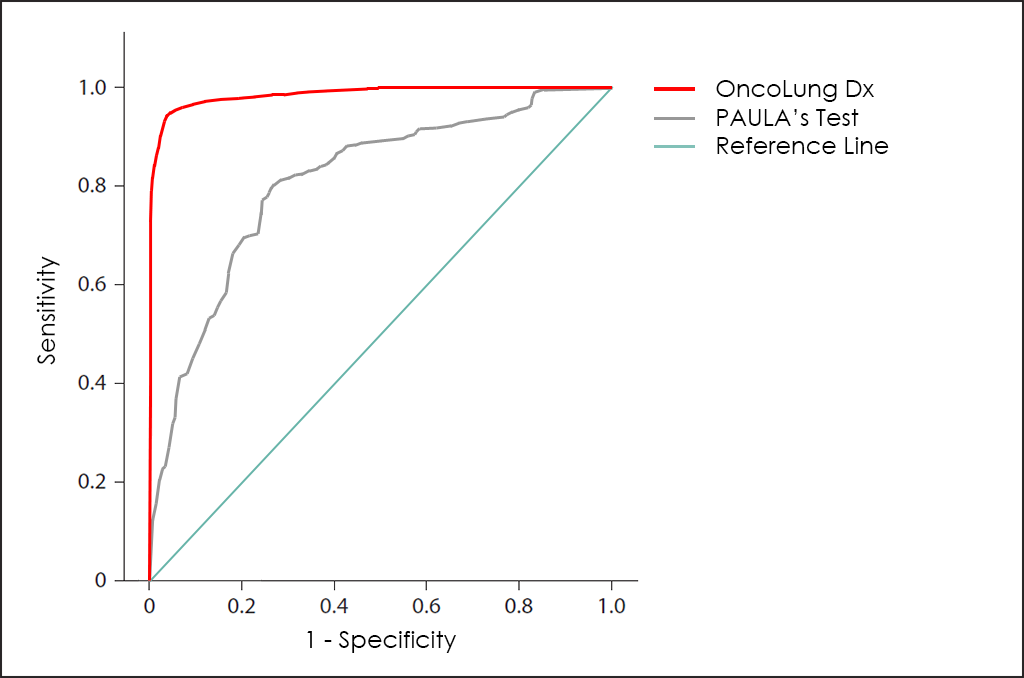
Besides, OncoLUNG Dx test is able to differentiate between Non-Small Cell Lung Cancer (NSCLC) and Small Cell Lung Cancer (SCLC) or Neuroendocrine ―while PAULA’s Test not―, and also determine the Histologycal Type and Subtype for NSCLC (Adenocarcinoma ―Large-Cell Lung Carcinoma―, and Squamous).
Uses and purposes for OncoLUNG Dx
OncoLUNG Dx test has been developed for:
- Aid in diagnostic assessments for high-risk patients (heavy smokers older than 55 years).
- Confirm or discard malignancy from results obtained previously with other tests, such as Computed Tomography (CT) Scan findings thanks higher Sensitivity and Specificity than imaging procedures.
- Help doctors predict the cancer’s behaviour and response to treatment, as well as a person’s chance of recovery.
- Guide treatment decisions (such as decide whether to add or immunotherapy after surgery and/or radiation therapy), therapy monitoring (doctors may use changes in the presence or amount of one or more Tumor Markers to assess how the cancer is responding to treatment) and predict or monitor for recurrence (looking for changes in the amount of a Tumor Marker may be part of their follow-up care plan and may help detect a recurrence sooner than other methods).
The Science Behind OncoLUNG Dx
Based on Publications
- Molina, R., Marrades R. M., Auge J. M., Escudero J. M., Vinolas N., Reguart N., Ramirez J., Filella X., Molins L. and Agusti A. (2016). “Assessment of a Combined Panel of Six Serum Tumor Markers for Lung Cancer.” Am J Respir Crit Care Med 193(4): 427-437. DOI: 10.1164/rccm.201404-0603OC
- Molina R., Filella X., Trapé J., Augé J. M., Barco A., Cañizares F., Colomer A., Fernandez A., Gaspar M. J., Martinez-Peinado A., Pérez L., Sánchez M., Escudero J. M. (2013). “Principales causas de falsos positivos en los resultados de marcadores tumorales en suero.” Sociedad Española de Bioquímica Clínica y Patología Molecular. Comisión de Marcadores Biológicos del Cáncer. PDF
Related Publications
- Black, W. C., I. F. Gareen, S. S. Soneji, J. D. Sicks, E. B. Keeler, D. R. Aberle, A. Naeim, T. R. Church, G. A. Silvestri, J. Gorelick, C. Gatsonis and T. National Lung Screening Trial Research (2014). “Cost-effectiveness of CT screening in the National Lung Screening Trial.” N Engl J Med 371(19): 1793-1802. DOI: 10.1056/NEJMoa1312548
- Chen, F., X. Y. Wang, X. H. Han, H. Wang and J. Qi (2015). “Diagnostic value of Cyfra21-1, SCC and CEA for differentiation of early-stage NSCLC from benign lung disease.” Int J Clin Exp Med 8(7): 11295-11300. PMCID: 26379938
- Doseeva, V., T. Colpitts, G. Gao, J. Woodcock and V. Knezevic (2015). “Performance of a multiplexed dual analyte immunoassay for the early detection of non-small cell lung cancer.” J Transl Med 13Cost-effectiveness of CT screening in the National Lung Screening Trial: 55. PMCID: PMC4335536
- Duffy, M. J. (2010). “Clinical Utility of Tumor Markers: What the Guidelines Recommend.” Journal of Laboratory Diagnostics 46(3): 281-291. PDF
- Duffy, M. J. and P. McGing (2010). Guidelines for the use of Tumour Markers. PDF
- Ferrigno, D., G. Buccheri and C. Giordano (2003). “Neuron-specific enolase is an effective tumour marker in non-small cell lung cancer (NSCLC).” Lung Cancer 41(3): 311-320. PMID: 12928122
- Fuentes, R., I. Bover, J. Rifa, V. Moreno, E. Canals, A. Marquez, E. Barreiro, J. Borras, R. Molina and P. Viladiu (2004). “Factors influencing survival in a prospective cohort of patients with non-small cell lung cancer: an updated assessment.” Clin Oncol (R Coll Radiol) 16(8): 583-584. “DOI: 10.1016/j.clon.2004.08.004“
- Gaspar, M. J., M. Diez, A. Rodriguez, T. Ratia, A. Martin Duce, M. Galvan, J. Granell and C. Coca (2003). “Clinical value of CEA and CA125 regarding relapse and metastasis in resectable non-small cell lung cancer.” Anticancer Res 23(4): 3427-3432.” PMID: 12926084
- Ghosh, I., D. Bhattacharjee, A. K. Das, G. Chakrabarti, A. Dasgupta and S. K. Dey (2013). “Diagnostic Role of Tumour Markers CEA, CA15-3, CA19-9 and CA125 in Lung Cancer.” Indian J Clin Biochem 28(1): 24-29. PMCID: PMC3547445
- Grenier, J., J. L. Pujol, F. Guilleux, J. P. Daures, H. Pujol and F. B. Michel (1994). “Cyfra 21-1, a new marker of lung cancer.” Nucl Med Biol 21(3): 471-476. PMID: 9296767
- Jett, J. R., L. J. Peek, L. Fredericks, W. Jewell, W. W. Pingleton and J. F. Robertson (2014). “Audit of the autoantibody test, EarlyCDT(R)-lung, in 1600 patients: an evaluation of its performance in routine clinical practice.” Lung Cancer 83(1): 51-55. DOI: 10.1016/j.lungcan.2013.10.008
- Korse, C. M., S. Holdenrieder, X. Y. Zhi, X. Zhang, L. Qiu, A. Geistanger, M. R. Lisy, B. Wehnl, D. van den Broek, J. M. Escudero, J. Standop, M. Hu and R. Molina (2015). “Multicenter evaluation of a new progastrin-releasing peptide (ProGRP) immunoassay across Europe and China.” Clin Chim Acta 438: 388-395. DOI: 10.1016/j.cca.2014.09.015
- Lam, S., P. Boyle, G. F. Healey, P. Maddison, L. Peek, A. Murray, C. J. Chapman, J. Allen, W. C. Wood, H. F. Sewell and J. F. Robertson (2011). “EarlyCDT-Lung: an immunobiomarker test as an aid to early detection of lung cancer.” Cancer Prev Res (Phila) 4(7): 1126-1134. DOI: 10.1158/1940-6207.CAPR-10-0328
- Manos, D., J. M. Seely, J. Taylor, J. Borgaonkar, H. C. Roberts and J. R. Mayo (2014). “The Lung Reporting and Data System (LU-RADS): a proposal for computed tomography screening.” Can Assoc Radiol J 65(2): 121-134. DOI: 10.1016/j.carj.2014.03.004
- Masuoka, T., Y. Matueda, H. Ookawa, K. Watanabe and S. Mimoto (1985). “[The measurement of SCC antigen in squamous cell carcinoma of the lung].” Gan No Rinsho 31(8): 914-918. PMID: 4032769
- Mayeux, R. (2004). “Biomarkers: potential uses and limitations.” NeuroRx 1(2): 182-188. DOI: 10.1602/neurorx.1.2.182
- Medicine, T. A. f. c. B. L. (2013). RECOMMENDATIONS AS A RESULT OF THE ACB NATIONAL AUDIT ON TUMOUR MARKER SERVICE PROVISION. PDF
- Molina, R. (2009). “ProGRP: A New Biomarker for Small Cell Lung Cancer.” EUROPEAN JOURNAL OF CLINICAL & MEDICAL ONCOLOGY 1(2): 25-32. DOI: 10.1016/j.clinbiochem.2004.05.007
- Molina, R., J. M. Auge, J. Alicarte, X. Filella, N. Vinolas and A. M. Ballesta (2004). “Pro-gastrin-releasing peptide in patients with benign and malignant diseases.” Tumour Biol 25(1-2): 56-61. DOI: 10.1159/000077724
- Molina, R., J. M. Auge, X. Bosch, J. M. Escudero, N. Vinolas, R. Marrades, J. Ramirez, E. Carcereny and X. Filella (2009). “Usefulness of serum tumor markers, including progastrin-releasing peptide, in patients with lung cancer: correlation with histology.” Tumour Biol 30(3): 121-129. DOI: 10.1159/000224628
- Molina, R., J. M. Auge, J. M. Escudero, R. Marrades, N. Vinolas, E. Carcereny, J. Ramirez and X. Filella (2008). “Mucins CA 125, CA 19.9, CA 15.3 and TAG-72.3 as tumor markers in patients with lung cancer: comparison with CYFRA 21-1, CEA, SCC and NSE.” Tumour Biol 29(6): 371-380. DOI: 10.1159/000181180
- Molina, R., J. M. Auge, X. Filella, N. Vinolas, J. Alicarte, J. M. Domingo and A. M. Ballesta (2005). “Pro-gastrin-releasing peptide (proGRP) in patients with benign and malignant diseases: comparison with CEA, SCC, CYFRA 21-1 and NSE in patients with lung cancer.” Anticancer Res 25(3A): 1773-1778. PMID: 16033098
- Molina, R., X. Bosch, J. M. Auge, X. Filella, J. M. Escudero, V. Molina, M. Sole and A. Lopez-Soto (2012). “Utility of serum tumor markers as an aid in the differential diagnosis of patients with clinical suspicion of cancer and in patients with cancer of unknown primary site.” Tumour Biol 33(2): 463-474. DOI: 10.1007/s13277-011-0275-1
- Molina, R., X. Filella and J. M. Auge (2004). “ProGRP: a new biomarker for small cell lung cancer.” Clin Biochem 37(7): 505-511. DOI: 10.1016/j.clinbiochem.2004.05.007
- Molina, R., X. Filella, J. M. Auge, R. Fuentes, I. Bover, J. Rifa, V. Moreno, E. Canals, N. Vinolas, A. Marquez, E. Barreiro, J. Borras and P. Viladiu (2003). “Tumor markers (CEA, CA 125, CYFRA 21-1, SCC and NSE) in patients with non-small cell lung cancer as an aid in histological diagnosis and prognosis. Comparison with the main clinical and pathological prognostic factors.” Tumour Biol 24(4): 209-218. DOI: 74432
- Molina, R., S. Holdenrieder, J. M. Auge, A. Schalhorn, R. Hatz and P. Stieber (2010). “Diagnostic relevance of circulating biomarkers in patients with lung cancer.” Cancer Biomark 6(3-4): 163-178. DOI: 10.3233/CBM-2009-0127
- Moro, D., D. Villemain, J. P. Vuillez, C. A. Delord and C. Brambilla (1995). “CEA, CYFRA21-1 and SCC in non-small cell lung cancer.” Lung Cancer 13(2): 169-176. PMID: 8581396
- Novakovic, S. (2004). “Tumor markers in clinical oncology.” Radiology and Oncology 38(2). PDF
- Oh, H. J., H. Y. Park, K. H. Kim, C. K. Park, H. J. Shin, J. H. Lim, Y. S. Kwon, I. J. Oh, Y. I. Kim, S. C. Lim, Y. C. Kim, S. H. Kim and M. G. Shin (2016). “Progastrin-releasing peptide as a diagnostic and therapeutic biomarker of small cell lung cancer.” J Thorac Dis 8(9): 2530-2537. DOI: 10.21037/jtd.2016.08.72
- Okamura, K., K. Takayama, M. Izumi, T. Harada, K. Furuyama and Y. Nakanishi (2013). “Diagnostic value of CEA and CYFRA 21-1 tumor markers in primary lung cancer.” Lung Cancer 80(1): 45-49. DOI: 10.1016/j.lungcan.2013.01.002
- Pan, Y. W., Z. G. Zhou, M. Wang, J. Q. Dong, K. P. Du, S. Li, Y. L. Liu, P. J. Lv and J. B. Gao (2016). “Combination of IL-6, IL-10, and MCP-1 with traditional serum tumor markers in lung cancer diagnosis and prognosis.” Genet Mol Res 15(4). DOI: 10.4238/gmr15048949
- Perez, E. O. and M. I. Aceituno Azaustre (2014). “Utilidad clínica de los marcadores tumorales.” Revista Médica de Jaén(4): 2-12. PDF
- Perkins, G. L., E. D. Slater, G. K. Sanders and J. G. Prichard (2003). “Serum Tumor Markers.” American Family Physician 68(6): 1075 – 1082. PDF
- Romero-Ventosa, E. Y., S. Blanco-Prieto, A. L. Gonzalez-Pineiro, F. J. Rodriguez-Berrocal, G. Pineiro-Corrales and M. Paez de la Cadena (2015). “Pretreatment levels of the serum biomarkers CEA, CYFRA 21-1, SCC and the soluble EGFR and its ligands EGF, TGF-alpha, HB-EGF in the prediction of outcome in erlotinib treated non-small-cell lung cancer patients.” Springerplus 4: 171. DOI: 10.1186/s40064-015-0891-0
- Schneider, J., M. Philipp, H. G. Velcovsky, H. Morr and N. Katz (2003). “Pro-gastrin-releasing peptide (ProGRP), neuron specific enolase (NSE), carcinoembryonic antigen (CEA) and cytokeratin 19-fragments (CYFRA 21-1) in patients with lung cancer in comparison to other lung diseases.” Anticancer Res 23(2A): 885-893. PMID: 12820318
- Sharma, S. (2009). “Tumor markers in clinical practice: General principles and guidelines.” Indian J Med Paediatr Oncol 30(1): 1-8. PMCID: PMC2902207
- Song, W. A., X. Liu, X. D. Tian, W. Wang, C. Y. Liang, T. Zhang, J. T. Guo, Y. H. Peng and N. K. Zhou (2011). “Utility of squamous cell carcinoma antigen, carcinoembryonic antigen, Cyfra 21-1 and neuron specific enolase in lung cancer diagnosis: a prospective study from China.” Chin Med J (Engl) 124(20): 3244-3248. PMID: 22088515
- Stieber, P., R. Hatz, S. Holdenrieder, R. Molina, M. Nap, J. von Pawel, A. Schalhorn, J. Schneider and K. Yamaguchi (2006). Practice Guidelines And Recommendations For Use Of Tumor Markers In The Clinic. National Academy of Clinical Biochemistry Guidelines for the use of tumor markers in Lung Cancer. PDF
- Sturgeon, C. (2002). “Practice guidelines for tumor marker use in the clinic.” Clin Chem 48(8): 1151-1159. PMID: 12142367
- Sturgeon, C. M., E. P. Diamandis, B. R. Hoffman, D. W. Chan, S. L. Ch’ng, E. Hammond, D. F. Hayes, L. A. Liotta, E. F. Petricoin, M. Schmitt, O. J. Semmes, G. Söletormos and E. van der Merwe (2009). National Academy of Clinical Biochemistry Laboratory Medicine Practice Guidelines for use of tumor markers in clinical practice: quality requirements. DOI: 10.1373/clinchem.2007.094144
- Sturgeon, C. M., M. J. Duffy, B. R. Hoffman, R. Lamerz, H. A. Fritsche, K. Gaarenstroom, J. M. G. Bonfrer, T. Ecke, H. B. Grossman, P. Hayes, R.-T. Hoffmann, S. P. Lerner, F. Lohe, J. Louhimo, I. Sawczuk, K. Taketa and E. P. Diamandis (2010). USE OF TUMOR MARKERS IN LIVER, BLADDER, CERVICAL, AND GASTRIC CANCERS, The National Academy of Clinical Biochemistry. PDF
- Trape, J., G. Gurt, J. Franquesa, J. Montesinos, A. Arnau, M. Sala, F. Sant, E. Casado, J. M. Ordeig, C. Bergos, F. Vida, P. Sort, A. Isava, M. Gonzalez and R. Molina (2015). “Diagnostic Accuracy of Tumor Markers CYFRA21-1 and CA125 in the Differential Diagnosis of Ascites.” Anticancer Res 35(10): 5655-5660. PMID: 26408739
- Trape, J., R. Molina, F. Sant, J. Montesinos, A. Arnau, J. Franquesa, R. Blavia, E. Martin, E. Marquilles, D. Perich, C. Perez, J. M. Roca, M. Domenech, J. Lopez and J. M. Badal (2012). “Diagnostic accuracy of tumour markers in serous effusions: a validation study.” Tumour Biol 33(5): 1661-1668. DOI: 10.1007/s13277-012-0422-3
- Trapé Pujol, J. and R. Molina Porto (2006). “Aspectos generales de los marcadores tumorales.” JANO 1620: 45-48. PDF
- Vinolas, N., R. Molina, M. C. Galan, F. Casas, M. A. Callejas, X. Filella, J. J. Grau, A. M. Ballesta and J. Estape (1998). “Tumor markers in response monitoring and prognosis of non-small cell lung cancer: preliminary report.” Anticancer Res 18(1B): 631-634. PMID: 9584045
- Yang, D. W., Y. Zhang, Q. Y. Hong, J. Hu, C. Li, B. S. Pan, Q. Wang, F. H. Ding, J. X. Ou, F. L. Liu, D. Zhang, J. B. Zhou, Y. L. Song and C. X. Bai (2015). “Role of a serum-based biomarker panel in the early diagnosis of lung cancer for a cohort of high-risk patients.” Cancer 121 Suppl 17: 3113-3121. DOI: 10.1002/cncr.29551
- Yang-Chun, F., F. Min, Z. Di and H. Yan-Chun (2016). “Retrospective Study to Determine Diagnostic Utility of 6 Commonly Used Lung Cancer Biomarkers Among Han and Uygur Population in Xinjiang Uygur Autonomous Region of People’s Republic of China.” Medicine (Baltimore) 95(18): e3568. DOI: 10.1097/MD.0000000000003568

